Subthalamic nucleus correlates of force adaptation
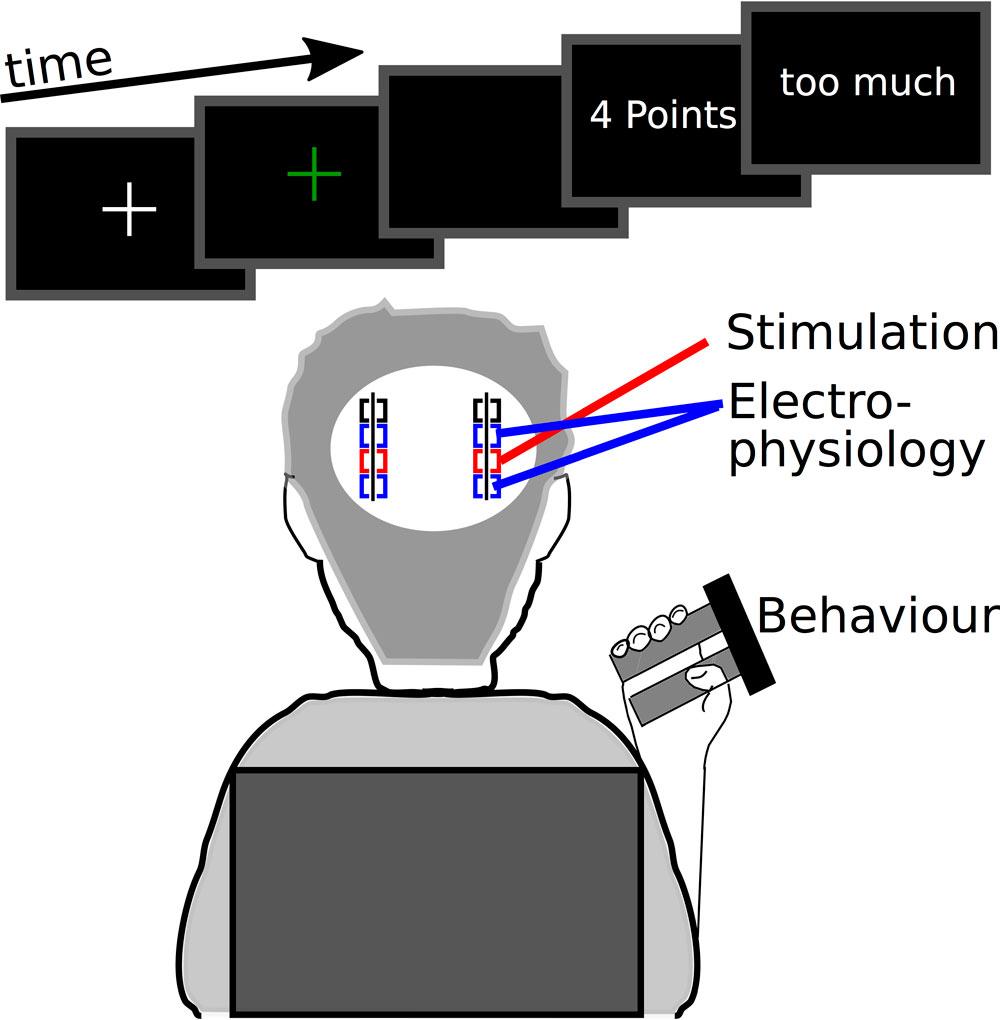
This code analyses behavioural data from a group of 16 Parkinson patients and 15 healthy control participants performing an action adaptation tasks, in which participants need to continuously adapt the applied force based on the feedback they receive. The first feedback ranges from 0 (worst) to 10 (best) points depending on the error between actual force and target force (Value-cue) and the second feedback indicates whether the force had been too low or too high (Direction-feedback). The main behavioural outcomes are measures of force production and force adaptation (folder 1, used for figure 1 in the published article). In patients local field potentials were recorded during the task and corresponding code is stored in folder 2 (figure 2&3). In 14 patients burst deep brain stimulation was applied during a second session. Its effects on behaviour and local field potentials are analysed with code from folder 3 and 4 (figures 4&5). The results have been published in a paper entitled ‘Neural underpinnings of action adaptation in the subthalamic nucleus’ by Herz et al.
The code has been tested on a MacBook Pro, macOS Mojave 10.14.6. All data were analysed in Matlab (2019a, requires a software license) and FieldTrip. Installation guides can be found on: https://matlab.mathworks.com/ and https://www.fieldtriptoolbox.org/download/. Run times of the different scripts is usually short (from < 1 minutes to ~ 5 minutes) for most analyses except for cluster-based permutation tests of linear mixed effects models, which take a few hours.
Example data is provided for the behavioural analysis for 2 healthy control (HC) participants. Of note, this is not the actual data from HC01 & HC02 from the study, but it allows testing the behavioural scripts if applicable (see below for instructions).
(i) Behavioral data:
Scripts:
‘CompareLevodopaDemographicsMVC’: this script compares demographics and the maximum voluntary contraction (MVC) between patients and HC, and tests the effect of levodopa on the Unified Parkinson’s Disease Rating Scale (UPDRS).
‘GetEvents’ (_PD & _HC): Imports the events file from PsychoPy using ‘ExtractData’ (& _HC) and saves it as a mat-file.
‘GetForce’ (_PD & _HC): Computes several variables reflecting force production and adaptation using several functions: ‘Forceparameters’ computes measures of force production and the time of peak force, and allows illustrating single trials, ‘Forces_within’ computes mean and standard error of mean (SEM) of single subject force traces, ’Stat_within’ computes several single subject correlations and measures of force adaptation, ‘Forces_across’ computes mean and SEM of group force traces, ‘stat_across’ computes group-mean trajectories of actual force and target force. The results are saved as a mat-file for subject-averaged data and a csv-file for single subject data.
‘Plot_Stats’ computes statistics of these measures of force production and adaptation and plots the results.
(ii) Local field potential data:
Scripts:
‘GetLFP_FirstLevel.m’: Loads data and applies preprocessing, time-frequency analysis and re-aligning of data using FieldTrip. It uses the custom-written functions.
‘MakeMontage_AllBipolar’ (which creates a bipolar montage from the monopolar data) and ‘EpochData_TF’ (which epochs the continuous data aligned to the feedback cue and peak force). The epoched spectra are saved.
‘GetLFP_SecondLevel_PlotSpectra.m’: Loads the spectra from first level analysis and plots the grand average as well as group-averaged beta, alpha (for feedback-aligned data) and gamma traces (for movement-aligned data).
‘GetLFP_SecondLevel_LME.m’: Loads the spectra from first level analysis and computes LME analyses with variables of interest using moving windows of single trial beta power.For cluster-based permutation tests (which take several hours) the function ‘PermTests_LME’ is used.
‘GetLFP_SecondLevel_controlLME.m’: Loads the spectra from first level analysis and computes control LME analyses: Effect of Value and Direction on Alpha power in the feedback period (where it showed an increase), effect of change in force and absolute change in force on Gamma power before peak force (where it showed an increase) and on Beta power after the Value feedback (where it showed a correlation with Value).
(iii) DBS effects on behaviour:
Scripts:
‘GetEvents_Stim’: Imports the events file from PsychoPy using ‘ExtractData’ and saves it as a mat-file.
‘GetForce_Stim’: Computes several variables reflecting force production and adaptation using several functions: ‘Forceparameters’ computes measures of force production and the time of peak force, and allows illustrating single trials, ‘Forces_within’ computes mean and standard error of mean (SEM) of single subject force traces, ‘Forces_across’ computes mean and SEM of group force traces. The results are saved as a mat-file for subject-averaged data and a csv-file for single subject data.
‘GetToS’ loads a file with the stimulation trace during the task, calls the function ‘ToS_DownsampleBinaryRemoveRamp’ (which downsamples the data to 1000Hz, makes stimulation binary (1 for ON, 0 for OFF) and removes the ramping so that only stimulation at effective intensities counts as stimulation) and loads the relevant behavioural data (change in force and absolute change in force). It then calls the functions ‘ToS_WindowedStim’ (which computes for each trial whether or not stimulation was given in any 100 ms moving windows for cue- and movement aligned data) and ‘ToS_Windowed_nexttrial’ (which computes change in force and absolute change in force for windows in which stimulation was applied vs. was not applied).The results are saved in a mat-file.
‘Plot_ToS’ loads this data, plots effects of stimulation on absolute change in force and change in force and provides statistics using cluster-based permutation tests (‘PermTests_ToS’). It also saves single trial behavioural data with a column stating whether DBS was applied in the critical time windows (which is used for the DBS effects on local field potentials analysis).
(iv) DBS effects on local field potentials:
Scripts:
‘GetLFP_FirstLevel_Stim.m’: Loads data and applies preprocessing, time-frequency analysis and re-aligning of data using FieldTrip analogously to the script described under (ii) except that it also detrends and demeans the data, applies a low-pass filter at 100 Hz and excludes noisy data points, which are then interpolated. The epoched spectra aligned to feedback and movement are saved.
‘GetLFP_FirstLevel_Stim_TrigOnset.m’: Same as above, but aligned to onset of stimulation bursts.
‘GetLFP_SecondLevel_Stim.m’: Loads data from the previous analysis, loads single trial data with info whether DBS was applied at critical time windows and plots these beta traces together with beta power off stimulation for time windows of interest. Cluster-based permutation tests are applied using the function ‘PermTests_ToS’. Grand-average spectra and
beta power irrespective of stimulation-timing are also plotted.
‘GetLFP_SecondLevel_TrigOnset.m’: Loads data from the previous _TrigOnset analysis and plots the group average.
(v) Downloaded scripts:
The following scripts were downloaded from mathworks.com: ‘computeCohen_d’ (measure of effect size), ‘jblill’ (filling significant clusters from permutation tests), ‘shadedErrorBar’ (illustrating mean and SEM).
(vi) Testing example data:
Two example datasets are provided (termed Kont01 & Kont02), which allow testing the behavioural force analysis. To do this the script GetForce_HC.m should be opened.
DirName and EventPath should be adjusted for the actual path.
Line 24: Should be changed to ‘for Subj=1:2’
Lines 103-108 should be commented, i.e. not used.
Setting plotforce to 1 (line 12) plots the single subject and group average force spectra.
Setting check to 1 (line 11) plots single trial force data. For this only use subject 1 or 2, not both (i.e. in line 24 use ‘for Subj=1’ or ‘for Subj=2’).
We welcome researchers wishing to reuse our data to contact the creators of datasets. If you are unfamiliar with analysing the type of data we are sharing, have questions about the acquisition methodology, need additional help understanding a file format, or are interested in collaborating with us, please get in touch via email. Our current members have email addresses on our main site. The corresponding author of an associated publication, or the first or last creator of the dataset are likely to be able to assist, but in case of uncertainty on who to contact, email Ben Micklem, Research Support Manager at the MRC BNDU.
Creative Commons Attribution-ShareAlike 4.0 International (CC BY-SA 4.0)
This is a human-readable summary of (and not a substitute for) the licence.
You are free to:
Share — copy and redistribute the material in any medium or format
Adapt — remix, transform, and build upon the material for any purpose, even commercially.
This licence is acceptable for Free Cultural Works. The licensor cannot revoke these freedoms as long as you follow the license terms. Under the following terms:
Attribution — You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
ShareAlike — If you remix, transform, or build upon the material, you must distribute your contributions under the same licence as the original.
No additional restrictions — You may not apply legal terms or technological measures that legally restrict others from doing anything the licence permits.

